flexrbs: Anatomy of Escherichia coli Ribosome Binding Sites
 description
description

 description
description
@article{Shultzaberger.Schneider2001,
author = "R. K. Shultzaberger
and R. E. Bucheimer
and K. E. Rudd
and T. D. Schneider",
title = "{Anatomy of \emph{Escherichia coli}
Ribosome Binding Sites}",
journal = "J. Mol. Biol.",
volume = "313",
pages = "215--228",
pmid = "11601857",
note = "\htmladdnormallink
{http://dx.doi.org/10.1006/jmbi.2001.5040}
{http://dx.doi.org/10.1006/jmbi.2001.5040}",
year = "2001"}
PDF Preprint copy.
Published 2001 October 16 at JMB, Abstract at Pubmed
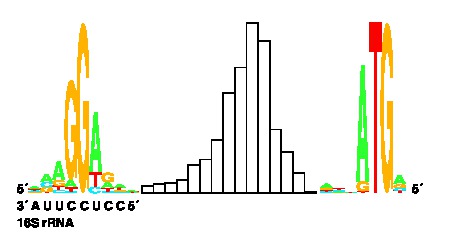
Summary of the flexible method: The basic observation is that the SD to Initiation Region (start codon and region around it, IR) distance is variable. One can, therefore, make a probability distribution, as shown above. One can compute the Shannon uncertainty of any distribution. This uncertainty remains after binding so it is to be subtracted from the sum of the other components. Furthermore, the ideas about individual information apply too and so one can build flexible sequence walker models. These work very well. The interesting thing is that one does not need to do any training to get these models. One starts from proven binding sites and gets the model directly. In contrast, training methods require that one provide examples of sequences that do not contain the site. However this is very difficult to obtain in general, so such training is probably contaminated with weak but functional sites. The information theory method avoids the problem.
The Delila Server allows you to try the model on the E. coli genome.
Supplementary data: Delila instructions for 569 verified ribosome binding sites from the EcoGene12 dabatase. These are N-terminal protein sequenced gene starts in E. coli that are uncleaved or have only the initiator methionine residue cleaved.
See also the companion paper, flexprom: Anatomy of Escherichia coli sigma 70 promoters
 as of 2005 Sep 17.
as of 2005 Sep 17.
We provide a table of data for
the rbseg12 model for the
U00096
E. coli K-12 MG1655 sequence
that contains the following information:
30 nucleotides upstream of gene start
Location of the gene start (Start)
Location of the ShineDalgarno (SD)
Orientation of the gene (Orient)
Strength of the SD (Ri(SD))
Distance between the SD and the ATG (Gap)
Total strength of RBS including ATG (Ri(total))
Here are the first two rows of the table:
*Sequence -30 to +2 ATG Start SD Orient Ri(SD) Gap Ri(total) cagataaaaattacagagtacacaacatccatg 190 175 1 5.47471 -15.0 5.86032
The first codon for every gene is the last three bases in the sequence. The SD coordinate corresponds to the central "G" = in the SD (refer back to the ribosome paper), and the spacing is the difference between this base and the first base of the start codon (usually an "A" in "ATG").
 as of 2005 Aug 23.
as of 2005 Aug 23.
CORRECTION: Under Materials and Methods page 225, third paragraph the reference to Blattner points to reference 37 instead of 33. For some reason we wrote that reference as 'Blattner et al. (1997)' and it got typeset incorrectly. Such errors never occur in LaTeX, which we use all the time; they occur frequently when people get involved, as apparently happened in this case. Unfortunately we missed the alteration at the proof stage. I, for one, am so used to the perfect referencing mechanism of LaTeX that I don't even think about checking such things anymore. But with humans involved, nothing is safe.
Other pointers:
![]()

Schneider Lab
origin: 2000 Jan 31
updated: 2019 Jun 21
![]()