
Schneider Lab
origin: 1999 May 21
updated: 2011 Aug 23
These are figures for:
@article{Wood.Wolf1999,
author = "T. I. Wood
and K. L. Griffith
and W. P. Fawcett
and K.-W. Jair
and T. D. Schneider
and R. E. Wolf",
title = "Interdependence of the Position and Orientation of
{SoxS} Binding Sites in the
Transcriptional Activation of the Class {I} Subset of
\emph{Escherichia coli}
Superoxide-Inducible Promoters",
journal = "Molec. Microb.",
volume = "34",
pages = "414-430",
year = "1999"}
"Interdependence of the Position and Orientation of SoxS Binding Sites in the Transcriptional Activation of the Class I Subset of Escherichia coli Superoxide-Inducible Promoters", T. I. Wood, K. L. Griffith, W. P. Fawcett, K.-W. Jair, T. D. Schneider, and R. E. Wolf, Molec. Microb., 34(3): 414-430. Pubmed entry.
Aligned SoxS binding sites, their individual information, sequence logo, and footprint data: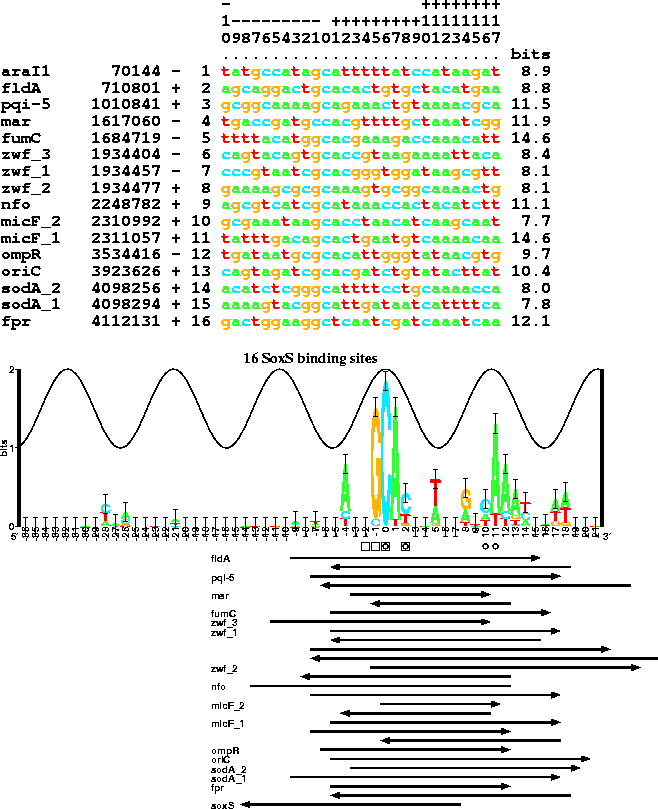
The sequences used to create this figure are defined by a set of Delila instructions. When Delila is run, the resulting small database is the book. The aligned listing is made by the Alist program.
Lister map showing sequence walkers at the locations of SoxS sites on various promoters relative to the estimated position of the -35 region: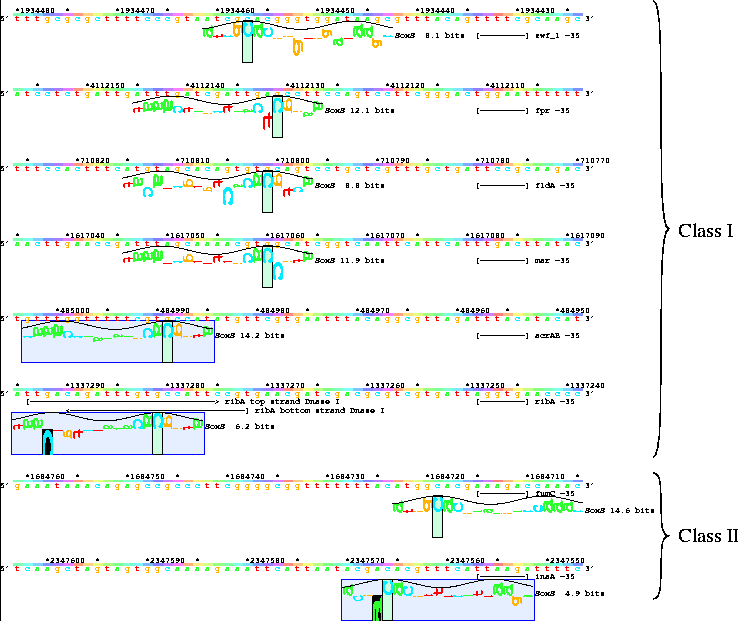
The colors of the lister wave are discussed on another page.
![]()

Schneider Lab
origin: 1999 May 21
updated: 2011 Aug 23
![]()