High Speed Parallel Molecular Nucleic Acid Sequencing
HIGH SPEED PARALLEL MOLECULAR NUCLEIC ACID SEQUENCING
PDF, thanks to www.freepatentsonline.com |

Image by Paul Theissan |
High Speed Parallel Molecular Nucleic Acid Sequencing
HIGH SPEED PARALLEL MOLECULAR NUCLEIC ACID SEQUENCING
PDF, thanks to www.freepatentsonline.com |

Image by Paul Theissan |
|
|
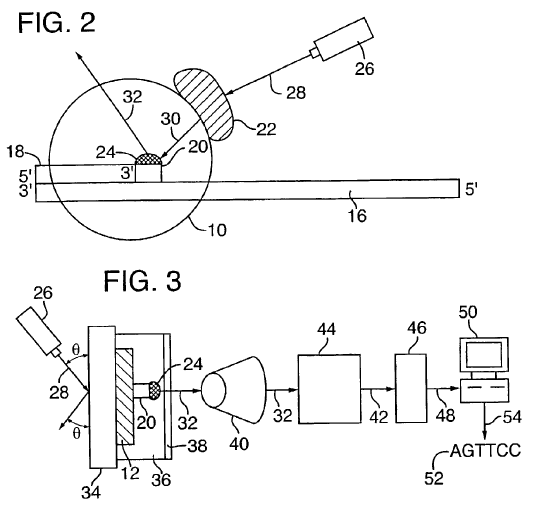
Fig 2:
To sequence a single DNA molecule (16),
a primer (18)
is annealed and a DNA polymerase (10) is bound.
Laser (26)
beam (28)
hits a Green Fluorescent Protein or other flurorophore (22)
attached to the polymerase.
Energy is transferred by
FRET (30)
to base (20)
which has a fluorescent compound attached (24).
The signal coming out (32)
can be seen in the microscope (Fig 3).
Fig 3: As the base (20) and the fluorescent dye (24) change, the output signal (32) changes. The signal is picked up by a microscope objective (40) and sent on to a computer (50) for processing into the final DNA sequence (52). |
Thomas D. Schneider, Denise Rubens (NCI)
Serial No. 60/151,580 filed 30 Aug 1999
Informal description of the Invention:
Using this technique the
sequence of single molecules of DNA or RNA can be read. The method uses a
microscope to observe changes in the fluorescence of individual nucleic acid
molecules being read by a DNA or RNA polymerase. The data are collected in
a computer in parallel, allowing many sequences to be determined from a tiny
volume. The technique could be used to read the sequences of mRNA directly
from cells, bypassing the current microarray technology. It may also be
fast enough to allow individual laboratories to read entire genomes with a
single device. This nanotechnology would have diagnostic uses by allowing
rapid identification of mutations that cause cancer and other genetic
diseases.
Formal description of the Invention: The present application describes a new method and apparatus for DNA sequencing called Two Dye Sequencing (TDS). This method employs engineered DNA polymerases which are labeled with a fluorophore such as Green Fluorescent Protein (GFP) and are combined with an annealed oligonucleotide primer in a chamber of a microscope field of view capable of detecting individual molecules. Four nucleotide triphosphates, each labeled on the base with a different fluorescent dye are introduced to the reaction. Light of a specific wavelength is used to excite the fluorophore on the polymerase, which in turn excites the neighboring fluorophore on the nucleotide by Forster Resonance Energy Transfer (FRET). As nucleotides are added to the primer, their spectral emissions provide sequence information of the DNA molecule.
See also Medusa Sequencing.
PCT Application (PDF) |
Federal Register Notifications |
![]()

Schneider Lab
origin: 2000 April 26
updated: 2013 Sep 11
![]()