 Molecular Computing Elements:
Molecular Computing Elements:
Description of the Invention:
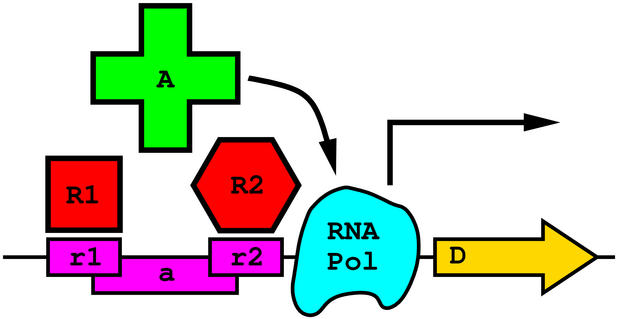
The basic idea of creating a molecular logic NOR gate is shown in the
figure at the top left. An RNA polymerase (cyan) can be activated
(arrows) by DNA binding protein A (green) which binds site
a
(purple).
Two other proteins R1 and R2 bind the DNA at sites
r1
and
r2
that overlap
with site
a
and interfere with binding of A. The resulting logic is a
NOR gate. All Boolean circuits can be built from NOR gates so it is
possible to build entire molecular computers with this gate.
The nice thing about such a logic gate is that is it very compact.
A more advanced form would use DNA binding proteins that stick strongly but which can be kicked off the DNA by GTP hydrolysis. I do not know if such things exist in nature, but it is likely that one could evolve them in the lab. The GTPase DNA binder can be triggered to come off the DNA by a GAP protein which could also bind DNA nearby. It is possible that such structures sitting on a meshwork of DNA origami could do lots of complex logic. Of course it would not be as fast as electronics but it could be grown and be massivly parallel (eventually).
Informal Description of the Invention:
A completely molecular
computer can be constructed using designed DNA and DNA binding proteins.
Several versions of the device are possible. In the simplest device,
bacteria or other cells are programmed with DNA sequences and DNA binding
proteins that execute logical Boolean operations. It is also possible to
construct molecular computers outside cells, but in this case special
attention must be paid to providing energy to run the computer. The
technology is based on the concept of a molecular flip-flop in which one or
more proteins compete for binding to binding sites that overlap. Because
only one protein can bind at a time but there are two ways for it to bind,
the method can also be used to double the sensitivity of diagnostic assays.
Formal Description of the Invention:
The present invention is a method and apparatus for molecular
computing which provides for molecular logic devices analogous to those
of electronic computers, such as flip-flops, AND gates, etc. Coupling
of the gates allows for molecular computing. The method allows data
storage, the transformation of binary information and signal readout.
Possible applications include encoding ``read only'' memory for
microscopic identifiers, digital control of gene expression, and
quantification of analytes. The computing elements also provide means
for complex regulation of gene expression.
|
European Patent No: 1057118
(PDF)
|
|
Granted August 10, 2004 United States Patent 6,774,222 at the US patent office United States Patent 6,774,222 at www.freepatentsonline.com |
 Molecular Flip-Flops Formed by Overlapping Fis Sites
is a paper describing the biology behind the idea.
Molecular Flip-Flops Formed by Overlapping Fis Sites
is a paper describing the biology behind the idea.
|
Article:
Molecular OS Gets Upgrade,
by Ivan Oransky in
The Scientist
(Volume 18, Issue 19, 38, Oct. 11, 2004),
describes our recently patented
method for molecular computing
 as of 2004 Oct 11.
:
as of 2004 Oct 11.
:
|

![]()

Schneider Lab
origin: 1998 June 16
updated:
2022 Jan 18: improve description
![]()