I thank Denise Rubens, Ilya Lyakhov, Herb Schneider, Natasha Klar, Bruce Shapiro, Richard Dawkins, Hugo Martinez and Karen Lewis for comments on the manuscript, and Frank Schmidt for pointing out that the atrophy should be first order.
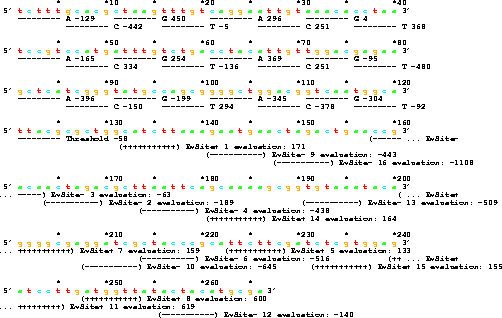
The organism has two parts, a weight matrix gene and a binding site region. The gene for the weight matrix covers bases 1 through 125. It consists of 6 segments 20 bases wide and one tolerance value 5 bases wide. Each segment contains sequence specifying the weights for the four nucleotides. For example, bases 1 to 5 contain tcttt. Translating this to binary gives 1101111111, which is the twos complement number for -129. This is the weight for A in the first position of the matrix. The 16 non-overlapping binding site locations were placed at random in the remaining portion of the genome. Evaluation by the weight matrix is indicated for each site. For example site 1, covering positions 132 to 137, catctt, is evaluated as -442 +296 -136 +251 +294 -92 = 171. Since this is larger than the threshold (-58), it is `recognized', and is marked with `+' signs. Evaluations to determine mistakes are for the first 256 positions on the genome. An extra 5 bases are added to the end, but not searched, to allow the sequence logos in Fig. 3 to have complete sequences available at all positions. Mutations are applied to all positions in the genome, so the binding sites and the weight matrix co-evolve. The figure was generated with programs ev, evd and lister. |
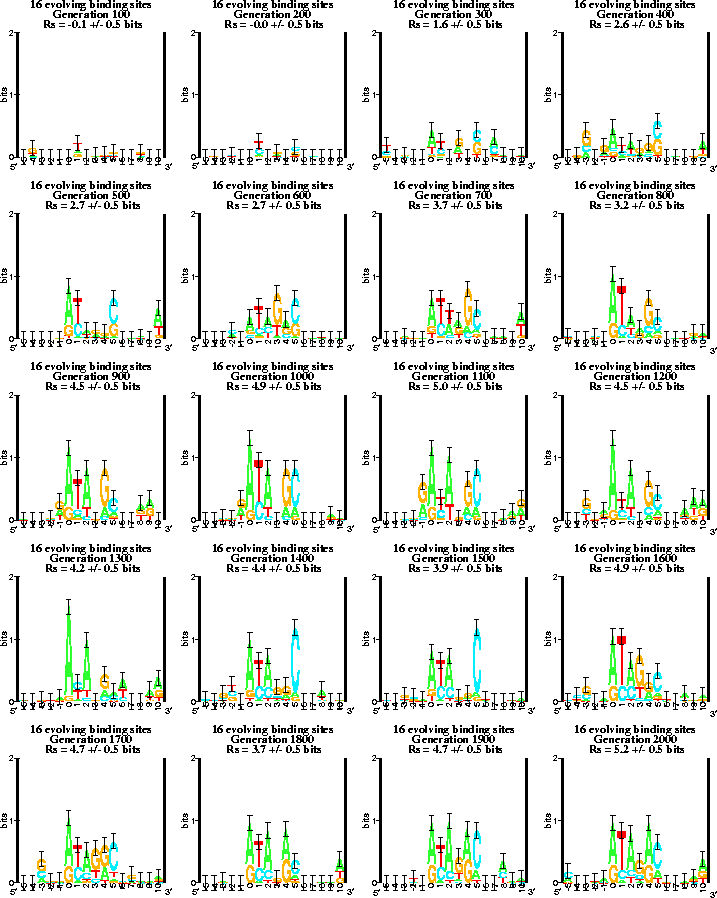
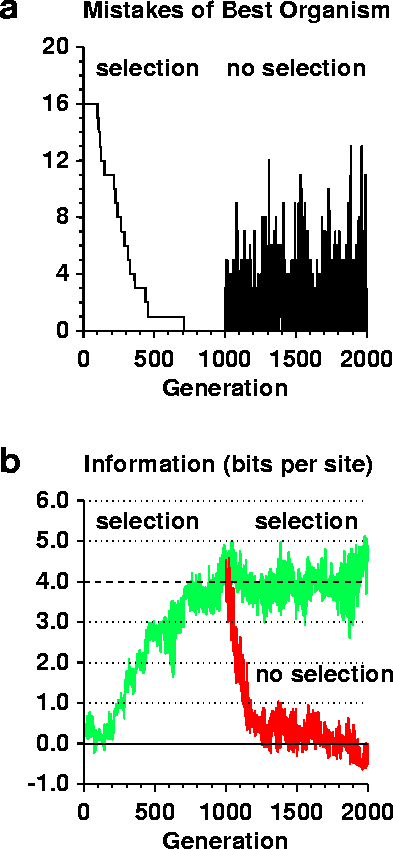
A sequence logo shows the information content at a set of binding sites by a set of stacks of letters [5]. The height of each stack is given in bits, and the sum of the heights is the total information content, Rsequence. Within each stack the relative heights of each letter are proportional to the frequency of that base at that position, f(b,l). Error bars indicate likely variation caused by the small sample size [4], as seen outside the sites, which cover positions 0 to 5. The complete movie is available at http://www.lecb.ncifcrf.gov//paper/ev/movie. |
![]()
![]()
![]()
Next: Bibliography
Up: Evolution of Biological Information
Previous: DISCUSSION
Tom Schneider
2001-11-07