
@article{Pluchino.Gottesman2015,
author = "K. M. Pluchino
and D. Esposito
and J. K. Moen
and M. D. Hall
and J. P. Madigan
and S. Shukla
and L. V. Procter
and V. E. Wall
and T. D. Schneider
and I. Pringle
and S. V. Ambudkar
and D. R. Gill
and S. C. Hyde
and M. M. Gottesman",
title = "{Identification of a Cryptic Bacterial Promoter in Mouse
(\emph{mdr1a}) P-Glycoprotein cDNA}",
journal = "PLoS One",
volume = "10",
pages = "e0136396",
pmid = "26309032",
year = "2015"}
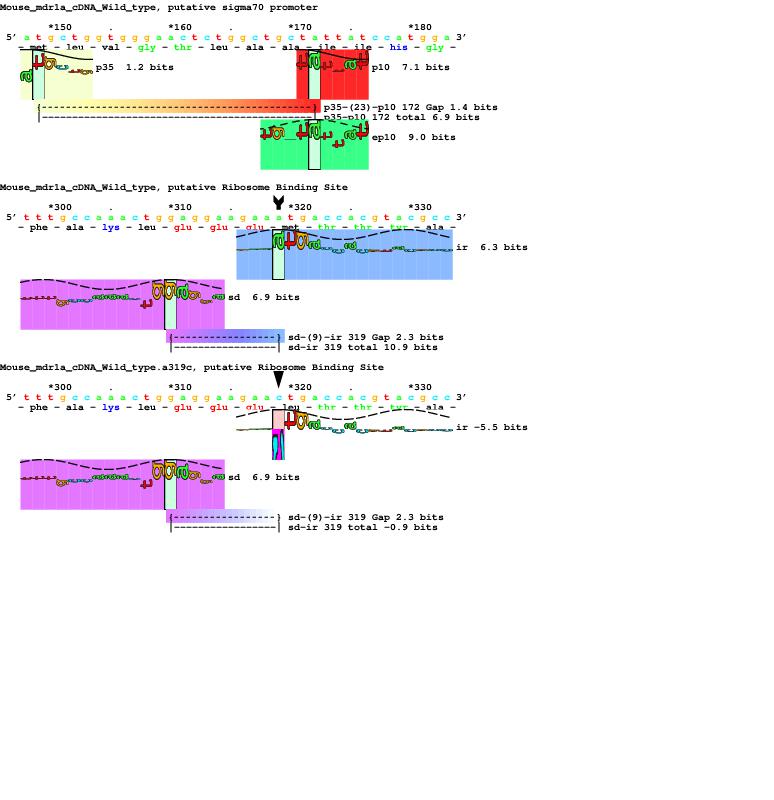
S1 Fig. Putative sigma 70 promoter and ribosome binding sites in mouse mdr1a cDNA.
The coordinate system begins at the first base of the mdr1a coding region. For the first two sequences, flexible individual information models for E. coli sigma70 (yellow, red rectangular petals) and extended -10 sigma70 (green) and ribosome binding sites (purple and blue) were scanned over the sequence and displayed using sequence walkers. The third sequence shows the effect of changing base 319 from an A to a C in M107L. This lowers the initiation codon from 6.3 bits to -5.5 bits, effectively removing the ribosome binding site. https://doi.org/10.1371/journal.pone.0136396.s001 (PDF)
Permanent link https://alum.mit.edu/www/toms/papers/Pluchino.Gottesman2015 points to the current web location.
![]()

Schneider Lab
origin: 2020 Feb 01
updated:
version = 1.00 of Pluchino.Gottesman2015.html 2020 Feb 01
![]()